Deworming Pill May Be Effective in Treating Liver Cancer
UCSF Researchers Use Computational Tools to Quickly Screen, Identify Drugs
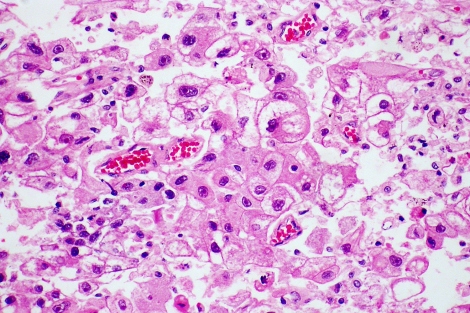
Hepatocellular carcinoma (HCC), a cancer associated with underlying liver disease and cirrhosis that often only becomes symptomatic when it is very advanced, is the second leading cause of cancer deaths around the world, and yet it has no effective treatment.
As with other conditions without treatments, the data that scientists need to understand and treat the disease may be sitting in plain view in databases that have barely been analyzed, says Atul Butte, MD, PhD, director of the Institute for Computational Health Sciences at UC San Francisco.
Bin Chen, PhD, a former postdoctoral scholar in Butte’s lab and now a faculty member in Pediatrics in the Institute for Computational Health Sciences, recently published a paper in Gastroenterology about using data-mining computational tools to identify a treatment for HCC.
Gene Expression and Drug Targets
Taking advantage of publicly available gene expression data, he first derived a molecular disease signature for HCC – looking at 274 genes that are either up or down regulated in cancerous liver tissues, but not in normal liver tissues.
Then, he looked for drugs that were known to target those genes and found, to his surprise, that a close cousin of a deworming pill, when used in combination with the standard care drug, was highly effective at killing cancerous liver tissue that had been engrafted into experimental mice.

“We found these disease genes were reversed after six weeks of treatment in a patient-derived tissue in mouse model,” Chen said, adding that the advantage of the approach he developed is that it targets a host of genes at the same time, rather than simply targeting a single mutation.
Chen said finding molecular signatures for diseases then looking for drugs that work against those signatures is a promising way of treating patients who may each have a different set of cancer mutations and might not respond to drugs that just targeted them one at a time.
“In this study, patients had similar gene expression profiles, but not identical,” Chen said. “In most cases, cancers have many mutations, and patients will relapse. Our approach might be used to control the disease, to make it a chronic condition rather than a lethal one.”
Head Start to a New Drug Candidate
Only a small number of drugs have been analyzed for their gene expression profiles, and so it was somewhat lucky that Chen found a hit with niclosamide. But even after discovering a match between the drug and the disease, his team of UCSF and Stanford University researchers was stymied. The drug, which was designed to kill parasites, did not work well in the mouse model, because it was not soluble.
It was only after Chen’s team stumbled upon a paper that described its close cousin, NEN – an ethanolamine salt that is soluble in human cells – that they made it work. When they combined NEN with sorafenib, the standard of care for advanced HCC, they found that the tumors stopped growing.
While NEN has been found safe to use against tapeworms in dogs and cats, scientists do not yet know how it would affect humans, or whether it would stop cancerous liver tissue from growing in humans.
But Chen said it is a promising drug candidate that could be developed at half or less than half the cost of a typical drug because of the head start he got by using advanced computational techniques to mine freely available data.
“Because our method is literally virtual, we can evaluate hundreds of drug candidates very quickly,” Chen said. “We looked at more than 1,000 drugs before discovering that the deworming pills were effective. This is a very efficient way to do drug discovery.”